Advanced Agricultural Biotechnology tools for Crop Improvement | informative 2023
Advanced Agricultural Biotechnology tools for Crop Improvement
Advanced Agricultural Biotechnology tools for Crop Improvement
Introduction:
For more than a century, plant breeding has played a significant role in raising crop productivity. Ongoing research activities and initiatives have been made to promote desirable traits like disease resistance, increased yield, resistance to abiotic stress, etc. in a single line or genotype. Crop cultivation is based on the objectives of novelty, stability, uniformity, and utility, which a breeder achieves by combining the use of conventional breeding with biotechnology tools; this emphasis of plant biotechnology enhances breeding for crop improvement. Thus, the quickest method to overcome the increasingly complex and overwhelming breeding process is through the integration of both plant breeding and biotechnology. To increase the likelihood of success, continuous varietal development through conventional breeding requires biotechnology. The two main biotechnological methods for improving crops are tissue culture and genetic engineering.
Biotechnology is more than genetic engineering when it comes to plant breeding because it solves issues across the board in the processing of agricultural products. This involves increasing and stabilizing yields, enhancing food’s nutritional value, such as the protein in pulses, etc., and enhancing resilience to pests, diseases, and abiotic conditions like drought and cold. Agricultural biotechnology, also known as agritech, is an area of agricultural science involving the use of scientific tools and techniques, including genetic engineering, molecular markers, molecular diagnostics, vaccines, and tissue culture, to modify living organisms: plants, animals, and microorganisms. Agricultural biotechnological features may receive enhanced cultivars in response to changing climatic conditions (Bernardo et al. 2008 & Varshney et al. 2011) and for the development of biotic and abiotic stress-resistant variations (Sharma et al. 2002).
Crop biotechnology is one aspect of agricultural biotechnology which has been greatly developed upon in recent times. Desired traits are exported from a particular species of crop to an entirely different species. These transgenic crops possess desirable characteristics in terms of flavor, color of flowers, growth rate, size of harvested products and resistance to diseases and pests. Farmers have manipulated plants and animals through selective breeding for ten thousands of years in order to create desired traits. Plant tissue culture, transgenic techniques, and molecular breeding methods are the three main components of biotechnology in crop breeding. Plant tissue culture is the process of growing plant cells or tissues in artificial media. It can be used for micropropagation, embryo rescue, protoplast culture, haploid creation, somaclonal hybridization, or somaclonal variants. Gene transfer from one organism to another is a significant use of biotechnology. This can be accomplished directly (via physical or chemical transfer) or indirectly (by agrobacterium-mediated gene transfer). The most common and widely used technique for crop improvement is molecular breeding, in which we use DNA markers and choosen varieties with better genetic diversity. The most common and widely used technique for crop improvement is molecular breeding, in which we use DNA markers and choose varieties with better genetic diversity. Recombinant DNA techniques, also known as genetic engineering or (more familiarly but less accurately) genetic modification, refer to the modification of an organism’s genetic make-up using transgenesis, in which DNA from one organism or cell (the transgene) is transferred to another without sexual reproduction. Genetically modified organisms (GMOs) are modified by the application of transgenesis or recombinant DNA technology, in which a transgene is incorporated into the host genome or a gene in the host, is modified to change its level of expression. The terms “GMO”, “transgenic organism” and “genetically engineered organism (GEO)” are often used interchangeably although they are not technically identical.
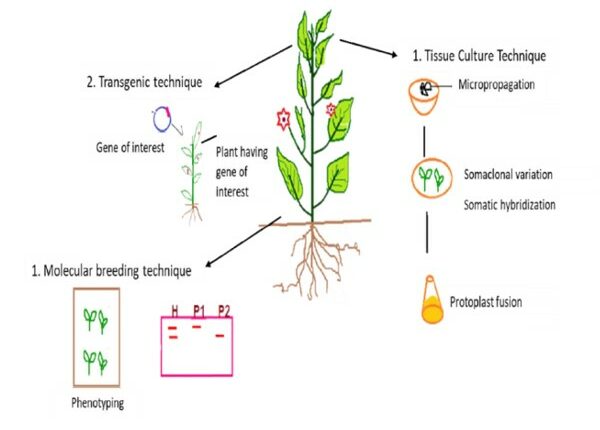
Molecular plant breeding is now helping to uncover genes and their activities, which may open up new directions for fundamental plant biology study (Moose and Mumm et al. 2008). In order to accelerate the rate of crop improvement, Watson et al. 2018, has recently concentrated on integrating speed breeding with other contemporary crop breeding methods, such as high-throughput genotyping, genome editing, and genomic selection. According to Croosa et al. 2017, Genomic selection (GS) facilitates the selection of superior genotypes in less time and speeds up the breeding cycle. The use of high throughput phenotyping, high throughput genotyping, genomic assisted breeding, and genome editing can improve crops more quickly than traditional methods. Figure1, demonstrating the several strategies of biotechnology for the improvement of cultivars.
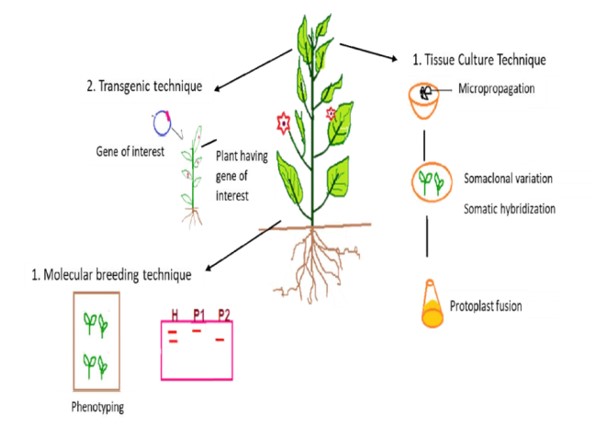
Figure1: Application of biotechnology tools in plant breeding practices (Source: Adlak et al. 2019)
Crop modification techniques:
- Traditional breeding: To increase crop quality and yield, traditional crossbreeding has been employed for generations. To produce a new variety with the desired qualities of the parents, two sexually compatible species must be crossed. For instance, because to the crossbreeding of its parents, the honey crisp apple has a particular texture and flavour. In conventional methods, pollen from one plant is applied to the female portion of another, resulting in a hybrid that has the genetic makeup of both parent plants. The plants that have the features that plant breeders want to pass on are chosen, and those plants are then bred again and again. Note that only within the same or closely related species can crossbreeding be used.
- Mutagenesis: Any organism’s DNA is subject to random mutations. Scientists can randomly introduce plant mutations to increase variety in crops. In the hopes of discovering the desired characteristic, mutagenesis uses radioactivity to cause random mutations. To introduce random mutations into DNA, scientists can employ radioactivity or compounds that cause mutations, such as EMS (ethylmethane sulfonate). Crop mutation is done in atomic gardens. A radioactive core is positioned in the middle of a circular garden and is lifted above the earth to radiate the crops nearby and cause mutations within a specific range. Ruby red grapefruits were created via mutagenesis induced by radiation.
- Polyploidy: In order to alter a crop’s fertility or size, polyploidy can be produced to change the number of chromosomes present in the crop. In most cases, organisms are diploid, or having two sets of chromosomes. However, that number of chromosomes can alter, either spontaneously or by the use of chemicals, leading to changes in fertility or crop size. In order to produce a sterile (seedless) watermelon with three sets of chromosomes, a 4-set chromosome watermelon is crossed with a 2-set chromosome watermelon.
- Protoplast fusion: For the purpose of transferring traits between species, protoplast fusion is the combining of cells or cell components. Using protoplast fusion, red cabbage, for instance, can inherit the feature of male sterility from radishes. The ability to produce hybrid crops thanks to male sterility.
- Cell/Tissue Culture & Micro propagation: To reproduce whole plants, it is possible to micropropagate small pieces of plant tissue or full structures, such as buds, in a lab setting. Micropropagation is particularly helpful for sustaining priceless plants, breeding species that would otherwise be challenging to breed (for example, many trees), accelerating plant breeding, and offering a wealth of plant material for research. Micropropagation is currently the foundation of a sizable commercial enterprise that involves hundreds of laboratories all over the world for species used in horticulture and crops. Micropropagation has the advantage of quick propagation, but it can also be used to produce planting material that is free of disease, especially when used in conjunction with diagnostic tools for disease detection. Some initiatives have been made to employ micropropagation in forestry more frequently. The increased multiplication rates offered by micropropagation when compared to vegetative propagation using cuttings.
- In vitro selection: The term “in vitro selection” refers to the selection of germplasm using a particular selection pressure on tissue culture in a lab setting. Numerous recent articles have demonstrated helpful links between in vitro responses and the production of desired field traits for crop plants, most frequently disease resistance. Positive results are also available for tolerance to metals, salt, low temperatures, and herbicides. The usefulness of in vitro selection is currently constrained by low correlations with field responses for the selection criteria of important general value in forest trees, including vigour, stem form, and wood quality. To test for disease resistance and tolerance to salt, frost, and drought, forestry programmes may find this method useful.
- Transgenics: In transgenics, a piece of DNA is inserted into the DNA of another organism in order to add a new gene or genes to the original organism. By introducing new genes into an organism’s genetic makeup, desired features are produced in a new variety. Before being put into the new organism, the DNA needs to be processed and packaged in a test tube. With biolistics, new genetic information can be added. The rainbow papaya, which has a gene altered to give it resistance to the papaya ringspot virus, is an illustration of a transgenic plant.
Only a few genetically modified crops are now offered for sale and consumption in the United States. Potatoes, apples, papaya, squash, canola, soybeans, corn, and canola have all received USDA approval. Arctic apples, a variety of GMO apples that don’t brown, reduce food waste, don’t require anti-browning treatments, and enhance flavour. With 10 million hectares of Bt cotton planted for the first time in 2011, the country’s Bt cotton production has surged, resulting in a 50% decrease in the need for insecticides. More than 15 million hectares of Bt cotton were sown in 2014 by Indian and Chinese farmers. Crops that have had their DNA altered through genetic engineering are referred to as genetically modified crops (or “GM crops” or “biotech crops”). For humans to produce vitamin A, beta carotene is necessary. The benefit of using golden rice is that it can be consumed in regions where vitamin A deficiency is frequent, hence preventing blindness. There are a few drawbacks of eating golden rice, including: the possibility of it contaminating wild rice through interbreeding. Hundreds of millions of people in developing nations depend mostly on rice as a staple meal. Thus, Golden Rice’s ability to provide -carotene may aid to reduce the chronic health issues brought on by vitamin A insufficiency (VAD).
Opportunities to use GM foods:
- Creating plants better resistant to weeds, pest and other diseases; such as corn.
- Bigger yields to create more efficient use of land, less uses of herbicides and other pesticides.
- Foods with better texture, flavor and nutritional value.
- Genetic engineering: Genetic engineering may be used to transfer a desired trait from an organism that is not sexually compatible with the host. The soil bacterium Agrobacterium tumefasciens is used as a vector in the most popular technique for genetic engineering plants. In order to infect the host plant, researchers must inject the desired gene or genes into the bacteria. Along with the infection, the desired genes are also passed on to the host. Dicot species like tomato and potato are the principal uses of this technique. Although the technique has lately been used effectively to transform wheat and other cereals, some crops, especially monocot species like rye and wheat, are not naturally amenable to transformation via tumefasciens. The target gene is coated on gold or tungsten particles and shot into the host at a high velocity using a “gene gun,” which is the most popular transformation method for these crops. It is evident that even distantly related genomes have common characteristics thanks to the creation of genomic maps that detail the precise location and sequences of genes. Through the rigorous study of a small number of genomes, comparative genomics aids in the comprehension of numerous genomes. For example, the mouse and malaria genomes serve as models for cattle and some of the diseases that impact them, and the rice genome sequence is beneficial for understanding the genomes of other cereals with which it shares traits according to its degree of relatedness. The majority of crops, animals, and diseases now have model species, and our understanding of their genomes is expanding quickly.
- Molecular markers: The inheritance of specific features can be identified using molecular markers, which are pieces of DNA. Hybridization- and PCR-based methods have been used to classify them (Gupta et al.1999). A great deal of progress has been made in the development and improvement of molecular techniques that make it simple to find markers of interest on a large scale. As a result, DNA markers are now widely and successfully used in human genetics, animal genetics and breeding, plant genetics and breeding, and germplasm characterization and management. The fundamental principle of plant breeding is the selection of suitable plant species, which includes assessing agronomic features, biotic and abiotic stress resistance/tolerance, and chemical reaction. A new branch of molecular breeding called marker assisted selection aids in the evaluation of qualities using molecular markers that are based on the banding pattern of associated DNA markers (Botstein et al. 1980). Restrictions fragment length polymorphism (RFLP), amplified fragment length polymorphism (AFLP), randomly amplified polymorphic DNA (RAPD), sequence-tagged sites (STS), expressed sequence tags (ESTs), simple sequence repeats (SSRs) or microsatellites, sequence-characterized amplified regions (SCARs), and single nucleotide polymorphisms are some of the DNA markers that have been developed and are used in plants (SNPs) (Nadeem et al. 2018 & Joshi et al. 1999). Table 1 highlights some of the molecular markers that have undergone comparative analysis and are utilized widely in molecular breeding methods.
Table 1: Comparative study of different types of molecular markers
| Characteristics | Restriction Fragment Length | Randomly Amplified Polymorphic DNA | Simple Sequence | Cleaved Amplified Polymorphic | Inter Simple Sequence Repeats | Expressed Sequence Tags | Single Nucleotide Polymorphism |
| Polymorphism
(RFLP) |
(RAPD) | Repeats (SSR) | Sequence (CAPS) | (ISSR) | (EST) | (SNP) | |
| Level of
polymorphism |
Medium | Very high | High | Moderate | High | High | High |
| Cost | Expensive | Cheap | Expensive | Cheap | Cheap | Costly then SSR | Variable |
| Allelism | Co-dominant | Dominant | Co-dominant | Mostly co-dominant | Dominant | Co-dominant | Co-dominant |
| Time | Time consuming | Quick working | Quick working | Quick | quick | Time Consuming | quick |
| Banding pattern | Locus specific | Multi locus | Locus specific | Locus specific | Multi locus | Locus specific | Locus specific |
| Probe / primer | Probe | Primer | primer | Primer | Primer | primer | Primer |
| DNA required (ng) | 10000 | 20 | 10-20 | 30-100 | 20-50 | 20-50 | 5-20 |
| Advantages | They are first DNA
marker discovered, Co-dominant and no need of prior sequencing |
Less DNA require,
easy to use and polymorphic |
Less DNA
required, highly reproductive |
Versatile, easily
scored and interpreted |
Highly polymorphic,
no need of prior sequencing |
Rapid and
inexpensive |
Widely distributed in
genome, co- dominant, highly reproductive |
| Disadvantage | Use of radioactive
probe & southern blotting step involved |
Low reproducibility,
dominant ,highly purified DNA is required |
High developing
cost, presence of more null allele |
Restriction
enzymes must be tested for polymorphisms |
Non-homology of
similar sized fragments, low reproducibility |
Lack of primer
specificity, labour oriented |
High developing cost |
| References | Botstein et al. 1980 | Williams et al., Welsh
et al. 1990 |
Tautz et al. 1984 | Konieczny et al.
1993 |
Gupta et al. 1994 &
Godwin et al.1997 |
Adams et al. 1991 &1993 | Michaels et al. 1998, Batley
et al. 2003 & Wiltshire et al. 2003 |
SNP’s are the most advanced molecular markers, but they are only available in well-equipped laboratories due to the need for technical know-how and expensive equipment and reagents. The use of molecular markers is expanding quickly across a variety of applications, including marker assisted foreground and background selection, gene pyramiding, QTL mapping, fine mapping, gene tagging, association mapping, TILLING and Eco-TILLING, among others. With the use of the markers, it is now possible to choose suitable plants even when they are still seedlings and to use those plants for hybridization in the same season. Selection is now independent of phenotype. Additionally, MAS makes it simple to combine horizontal resistance with vertical resistance, which is quite challenging based on illness studies, and to pyramid oligogenic resistance. A number of plant genes have been positionally cloned using closely related molecular markers. In order to create variants with improved adaptation, novel breeding strategies like genomic selection (GS) and genomic assisted breeding (GAB) Scheme have been sparked by the discovery of molecular markers.
- Genomic Assisted Breeding and High Throughput Genotyping: The development of new tools in genomics research by bioinformatics is gradually improving the effectiveness and precision of crop improvement. In the end, whole genome selection and genotype improvement for any trait may be accomplished in silico by the breeder using relative values of targeted alleles at a given location in a segregating population. In order to understand how crop species with adaptive features respond to stress or to locate underlying genes, alleles, or quantitative trait loci, genomics is the most effective approach available. The most effective methods for improving crop plants’ ability to adapt to biotic and abiotic stresses are molecular breeding techniques, and recent developments in high throughput genotyping, sequencing, and phenotyping platforms (phenomics) have changed molecular breeding into genomics assisted breeding (GAB). Marker assisted selection (MAS) and genomic selection are the two methods for genomically assisted breeding that are most frequently utilised (GS) (Qian et al. 2018 & Carley et al. 2017). The SNP array, a comparatively inexpensive and automated genotyping technique, is used for high throughput genotyping. Genome-wide association studies (GWAS), linkage map construction, genomic selection, population structure analysis, and gene mapping are only a few of the genetic studies of crops that have extensively employed it. Currently, hybridization-based SNP arrays and different NGS-enabled genotyping methods including GBS and GS are the most widely used high throughput genotyping technologies. For high-throughput genotyping of crops, SNP array has several advantages over traditional marker based genotyping and NGS approaches. Some of the points includes, SNP array data is relatively easy to analyze compared to data generated using NGS-based methods, labor intensive NGS library preparation and bioinformatics data analysis investment for accurate SNP calling. The use of whole genome sequencing with high coverage and the upgrading of the markers on the SNP array are two steps done to lessen these constraints. Widely used platforms from Illumina and Affymetrix are for SNP marker selection in array building. SNP depth, SNP kinds, SNP frequency, extra changes within the target SNP probe sequence, and probe sequence parameters should all be taken into consideration when choosing an SNP array. The possibility of using natural populations in a genome-wide association study (GWAS) has been made possible by the availability of high-density SNP markers. GWAS could circumvent the limitations of traditional linkage mapping and offer a potent alternate approach for analysing complex variables. By examining characteristics and high-density marker scores within an artificially produced population at the wholegenome level, genomic selection (GS) forecasts genomic projected breeding values of lines. GS is an additional viable breeding technique for quickly enhancing complicated features. GS has been shown to be superior to marker assisted recurrent selection for improving complex traits in crops, but still being more expensive, because it can successfully avoid problems related to the number of QTL that govern a trait.
Future prospects:
For determining evolutionary links and deciphering the functions of certain genes, genomic sequences of organisms are vitally significant. The breeding process will be accelerated by the discovery of the genes and molecular markers underlying agronomic features, which will result in superior varieties with increased yield and quality, tolerance to unfavourable environmental circumstances, and resistance to diseases. DNA sequencing is a useful assay, and as it becomes faster and less expensive, it will find more and more uses in plant breeding. With the advent of next-generation sequencing, it is now much easier and less expensive to analyse the mutations that appear in an organism’s entire genome. Crop genome organisation and structure can be learned a lot through crop sequencing. It creates an abundance of possibilities for life sciences research, including evolutionary biology, developmental biology, biochemistry, genetics, and molecular biology. A second technical revolution in DNA sequencing has recently been underway in the field of agricultural sciences. Despite the fact that agricultural yield and output have improved greatly as a result of conventional breeding procedures, new strategies are needed to further enhance crop productivity in order to fulfil the world’s expanding food demand. The genome editing tool based on Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas9 (CRISPR-associated protein9) has shown significant potential for quickly resolving new problems in agriculture. Recently, Haque et al. 2018, reported the potential of CRISPR/Cas9 for enhancing tropical climate-adapted crops to become more resilient to new pests and abiotic challenges. Any organism, including plants, can have its genome sequence precisely modified to produce the desired feature. Numerous strategies have recently been tested to enhance plant transformation using CRISPR/Cas9, including optimization of the promoters to drive and express Cas9 and use of various fluorescent reporters and selection markers (Wang et al. 2015, Yan et al.2015, Kaur et al. 2018 & Gao 2018). The CRISPR/Cas gene-editing technique may produce heritable, targeted alterations and can also produce transgene-free plants, which allays worries about the presence of foreign DNA sequences. Rice is the crop that has been studied the most, followed by maize, tomato, potato, wheat, barley, and other important crops. The ongoing development of biotechnology methods will undoubtedly aid in increasing crop yield while maintaining sustainability.
References:
- Adams MD, Kelley JM, Gocayne JD Dubnick M, Polymeropoulous MH, Xiao H, Merril CR, Andrew WU, Olde B, and Venter JC (1991). Complementary DNA sequencing: expressed sequence tags and human genome project. Science, 252:1651- 1656.
- Adams MD, Soares MB, Kerlavage AR, Fields C, Venter JC (1993). Rapid cDNA sequencing (expressed sequence tags) from a directionally cloned human infant brain cDNA library. Nature Genetics, 4:373-380.
- Adlak T, Tiwari S, Tripathi MK, Gupta N, Sahu VK, Bhawar P and Kandalkar VS (2019). Biotechnology: An Advanced Tool for Crop Improvement Current Journal of Applied Science and Technology 33(1): 1-11, 46931:2457-1024.
- Batley J, Barker G, ‘Sullivan HS, Edwards KJ and Edwards D (2003). Mining for single nucleotide polymorphisms and insertions/deletions in maize expressed sequence tag data. Plant Physiology, 132(1):84-91.
- Bernardo R (2008). Molecular markers and selection for complex traits in plants: learning from the last 20 years Crop Science, 48(5): 1649-1664.
- Botstein D, White RL, Skolnick M, Davis RW (1980). Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics, 32(3):314-31.
- Carley CAS, Coombs JJ, Douches DS, Bethke PC, Palta JP, Novy RG, Endelman JB (2017). Automated tetraploid genotype calling by hierarchical clustering. Theoretical & Applied Genetics, 130: 717-726.
- Crossa J, Pérez-Rodríguez P, Cuevas J Montesinos-López O, Jarquín D, de Los Campos G, Burgueño J, González-Camacho JM , Pérez-Elizalde S, Beyene Y, Dreisigacker S, Singh R, Zhang X, Gowda M, Roorkiwal M, Rutkoski J, Varshney RK (2017). Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Science, 22: 961-975.
- Gao C. (2018). The future of CRISPR technologies in agriculture. Nature Reviews Molecular Cell Biology. 19:275-276.
- Godwin ID, Aitken EAB and Smith LW (1997). Application of inter simple sequence repeat (ISSR) markers to plant genetics. Electrophoresis, 18:1524-1528.
- Gupta M, Chyi YS, Romero-Severson, J and Owen JL (1994). Amplification of DNA markers from evolutionarily diverse genomes using single primers of simple sequence repeats’, Theoretical and Applied Genetics, 89: 998-1006.
- Gupta PK, Varshney RK, Sharma PC and Ramesh B (1999). Molecular markers and their applications in wheat breeding. Plant Breeding, 118(5):369-390.
- Wang ZP, Xing XL, Dong L, Zhang HY, Han CY, Wang XC & Chen QJ (2015). Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biology, 16: 144.
- Haque E, Taniguchi H, Hassan MM, Bhowmik P, Karim MR, Smiech M, Zhao K, Mahfuzur R and Islam T* (2018). Application of CRISPR/Cas9 Genome Editing Technology for the Improvement of Crops Cultivated in Tropical Climates: Recent Progress, Prospects and Challenges. Frontiers in Plant Science, 9:617.
- Joshi SP, Ranjekar PK and Gupta VS (1999). Molecular markers in plant genome analysis. Current Science, 77:230- 240.
- Kaur N, Alok A Shivani, Kaur N, Pandey P, Awasthi P Tiwari S (2018). CRISPR/Cas9-mediated efficient editing in phytoene desaturase (PDS) demonstrates precise manipulation in banana cv. rasthali genome. Functional Integrative Genomics, 18:89-99.
- Konieczny A, Ausubel FM (1993). A procedure for mapping arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant Journal, 4:403-410.
- Michaels SD and Amasino RM (1998). A robust method for detecting single-nucleotide changes as polymorphic markers by PCR. Plant Journal. 14:381-385.
- Moose S and Mumm R (2008). Molecular Plant Breeding as the Foundation for 21st Century Crop Improvement. Plantphysiology, 147: 969-77.
- Nadeem MA, Nawaz MA Shahid MQ, Dogan Y, Comertpay G, Yıldız M, Hatipoglu R, Ahmad F, Alsaleh A, Labhane N, Ozkan H, Chung G, Baloch SF (2018). DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnology and Biotechnological Equipment, 32 (2):261-285.
- Qian Y, Xiping Y, Peng Z, Xu L and Wang J (2018). Development and Applications of a High Throughput Genotyping Tool for Polyploid Crops: Single Nucleotide Polymorphism (SNP) Array. Frontiers in Plant Science, 9:2018, https://doi.org/10.3389/fpls.2018.00104.
- Sharma HC, Crouch JH, Sharma KK, Seetharama N and Hash CT (2002). Applications of biotechnology for crop improvement:prospects and constraints. Plant Science. 163 (3):381-395.
- Tautz D, Renz M. (1984). Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Research, 12:4127-4138.
- Varshney RK, Bansal KC, Aggarwal PK, Datta SK and Craufurd PQ (2011). Agricultural biotechnology for crop improvement in a variable climate: hope or hype? Trends in plant science, 16 (7):363-371.
- Watson A, Ghosh S, Williams MJ, Cuddy WS, Simmonds J, Rey MD, Hatta, MA Md, Hinchliffe A, Steed A, Reynolds D, Adamski NM, Breakspear A, Korolev A, Rayner T, Dixon LE, Riaz A, Martin W, Ryan M, Edwards D, Batley J, Raman H, Carter J, Rogers C, Domoney C, Moore G, Harwood W, Nicholson P, Dieters M J , DeLacy IH, Zhou J, Uauy C, Boden SA, Park RF, Wulff BBH , Hickey LT (2018). Speed breeding is a powerful tool to accelerate crop research and breeding. Nature Plants, 4:23-29.
- Welsh J, McClelland M (1990). Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Research, 18:7213- 7218.
- Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research, 18:6531- 6535.
- Wiltshire T, Pletcher M T, Batalov S, Barnes S W, Tarantino L M (2003). Genome-wide single-nucleotide polymorphism analysis defines haplotype patterns in mouse. Proceedings of National Academy Sciences, 100(6):3380-3385.
- Yan L, Wei S, Wu Y, Hu R, Li H, Yang W and Xie Q (2015). High-Efficiency genome editing in Arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Molecular Plant, 8: 1820-1823.
Use of Advanced Agricultural Biotechnology tools for Crop Improvement
Sushma Kumari Pawar1, Swati Sharma1, Chandra Nath Mishra*1, Vijeta Sagwal1, Nishu Raghav1, Ruchika Sharma1, Kiran Devi1 and Joginder Singh Duhan2
1ICAR-Indian Institute of Wheat and Barley Research, Karnal, Haryana-132001
2Chaudhary Devi Lal University, Sirsa, Haryana-125055
=======================================================================================
Note
despite of that if there is any query please feel free contact us or you can join our pro plan with rs 500 Per Year, for latest updates please visit our modern kheti website www.modernkheti.com join us on Whatsapp and Telegram 9814388969. https://t.me/modernkhetichanel


Comments are closed.